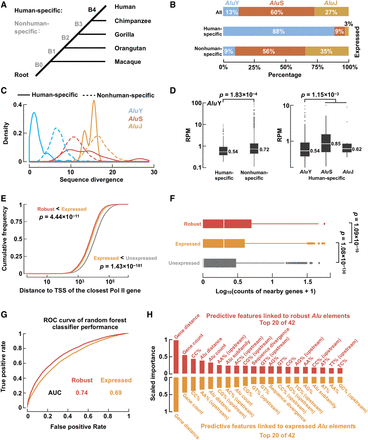
Expressed Alu elements show distinct genomic context and sequence features. (A) The primate phylogeny. B4 denotes the human-specific branch, and B0–B3 denote nonhuman-specific branches. (B) The AluY subfamily accounted for a large proportion of human-specific Alu elements. (C) Sequence divergence distributions of human-specific (solid lines) and nonhuman-specific (dashed lines) Alu elements in each Alu subfamily. Note that human-specific AluY elements show lower sequence divergence than nonhuman-specific AluY elements. (D) Human-specific AluY elements showed lower expression levels (measured by RPM) than nonhuman-specific AluY elements (left) and human-specific AluS/J elements (right). Wilcoxon rank-sum test P-values are shown. (E) Robustly expressed (red) and expressed (orange) Alu elements are closer to the TSS of Pol II–transcribed genes than are unexpressed (gray) Alu elements. Wilcoxon rank-sum test P-values are shown. (F) Robustly expressed (red) and expressed (orange) Alu elements are more likely to be located in gene-rich regions than are unexpressed (gray) Alu elements. Wilcoxon rank-sum test P-values are shown. (G) Receiver operating characteristic (ROC) curve of random forest classifiers for distinguishing robustly expressed or expressed against unexpressed Alu elements using genomic context and sequence features. (AUC) Area under the curve. (H) The top 20 most important features of the random forest classifiers for distinguishing robustly expressed (red bars) or expressed (orange bars) Alu elements against unexpressed Alu elements, ordered by feature importance.











