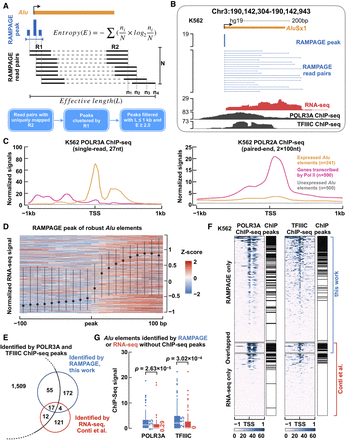
Genome-wide identification of expressed Alu elements using RAMPAGE data. (A) A computational pipeline to call RAMPAGE peaks and annotate expressed Alu elements. Paired-end RAMPAGE reads (R1 and R2, black; N, read pairs in total) with uniquely mapping R2 reads were first clustered to call peaks at the 5′-end of R1, and the resulting peaks (blue bars) were further filtered with entropy (E) and effective length (L) to annotate expressed Alu elements (orange bar) (for more details, see Supplemental Methods). (B) An expressed AluSx1 element in an intergenic region. RAMPAGE peak marks the TSS of the AluSx1 element precisely, with RAMPAGE read pairs linking the TSS to downstream positions. RNA-seq and ChIP-seq data sets of POLR3A and TFIIIC further confirm the expression of this AluSx1 element. (C) POLR3A (left) and POLR2A (right) ChIP-seq signals are shown with respect to the TSS of expressed Alu elements (orange line), protein-coding and lncRNA genes transcribed by Pol II (pink line), and unexpressed Alu elements (gray line). The signals were averaged over all genes in each set. (D) RNA-seq signal in the ±100-bp window (in 10-nt bins) centered on the TSSs of robustly expressed Alu elements identified by RAMPAGE (identified in three or more biosamples). To avoid overlap with the TSSs of Pol II–transcribed genes, only intronic and intergenic Alu elements were included. Each row of the heatmap corresponds to one such Alu element in one biosample with 10 or more nonzero RNA-seq signal bins, and the dot and bars correspond to the median and first and third quartiles of all Alu elements. (E) Only 84 of the 1593 Alu elements that overlap POLR3A and TFIIIC ChIP-seq peaks (Moqtaderi et al. 2010) are expressed according to RAMPAGE or RNA-seq data in K562 cells. On the other hand, 297 expressed Alu elements (by RAMPAGE or RNA-seq; four by both) do not overlap POLR3A and TFIIIC peaks. (F) Heatmap of normalized read densities for POLR3A and TFIIIC ChIP-seq data around the TSSs of expressed Alu elements identified by RAMPAGE, RNA-seq (Conti et al. 2015), or both techniques in K562 cells. ChIP-seq peaks (Moqtaderi et al. 2010) are labeled on the right of the corresponding heatmaps. (G) Among the 219 expressed Alu elements (by RAMPAGE or RNA-seq) that do not overlap POLR3A or TFIIIC peaks, the Alu elements identified by RAMPAGE show significantly higher POLR3A and TFIIIC signals than the Alu elements defined by RNA-seq. Wilcoxon rank-sum test P-values are shown.











