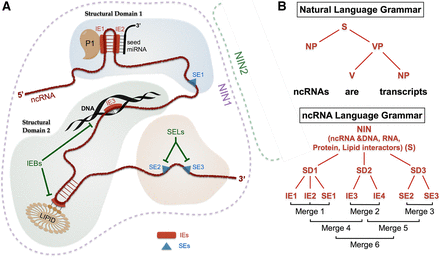
The analogy between the natural language grammar and the ncRNA structure grammar. (A) The elements of the ncRNA language grammar. A schematic view of IEs, SEs, structural domains of a ncRNA, and the noncoding RNA interactor network (NIN) composed by the ncRNA, interactor RNAs (such as miRNAs), interactor proteins (such as P1), interactor DNA elements, and interactor lipids (such as PIP3). Each ncRNA can contain multiple structural domains (here we show three for simplicity): one or multiple IEs and/or one or multiple SEs. These elements can be targeted by IE-blockers (IEBs), which release a specific interactor molecule (either DNAs, RNAs, proteins or lipids), and SE-lockers (SELs), which lock the lncRNA structure in a specific conformation favoring specific interactions with multiple molecules. A new type of therapy based on the correction of various interconnected genetic alterations that occur in the complex NINs by targeting the IEs can be envisaged, because these are docking sites for multiple types of molecules (DNA, RNA, proteins, and lipids) and/or the SEs, that directly affects the conformation of a lncRNA and indirectly the functional interactions with interactor molecules. (B) A comparison between the natural language and ncRNA language grammars. The various elements of the ncRNA language grammar are assembled under a “merge” (combine) function: The IEs and SEs from a lncRNA are combined during evolution by multiple rounds of merge (here, steps 1–6). (S) Subject; (NP) noun phrase; (VP) verb phrase; (V) verb.











