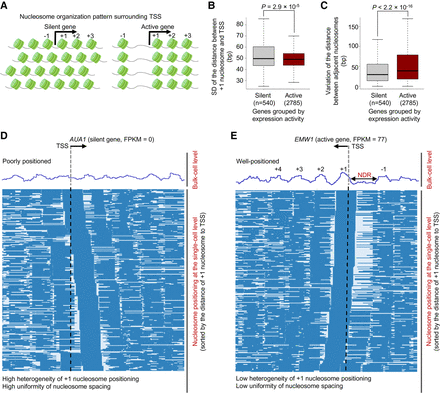
Differential nucleosome organization principles for silent and active genes. (A) Previous studies revealed nucleosome organization patterns surrounding the TSS of silent (left) and active (right) genes (Lai et al. 2018). Nucleosome positioning in promoter regions of silent genes showed large variation among cells but was highly uniformly spaced within each cell. In contrast, nucleosome positioning surrounding the TSS of active genes showed little variation among cells but relatively nonuniform spacing within each cell. (B) Heterogeneity of nucleosome positioning for silent (FPKM = 0) and active (FPKM > 50) genes. The heterogeneity of nucleosome positioning was measured by the standard deviation (SD) of the distances between +1 nucleosomes and the TSS. The P-value was calculated by the Wilcoxon rank-sum test. (C) Uniformity of nucleosome spacing for silent (FPKM = 0) and active (FPKM > 50) genes. See Methods for the definition of uniformity. The P-value was calculated by the Wilcoxon rank-sum test. (D) Long-range nucleosome positioning patterns for the silent AUA1 across different cells. Each row represents a cell, and nucleosome is labeled as blue bar. (E) Long-range nucleosome positioning patterns for the actively transcribed gene EMW1 across different cells.











