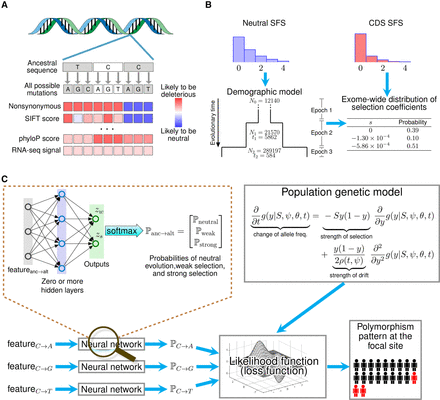
Overview of LASSIE. (A) For each potential protein-coding mutation, we collected 33 genomic features likely to be informative about natural selection, including variant categories, protein and nucleotide conservation scores, and RNA-seq signals (Supplemental Table S1). (B) A three-epoch demographic model was fitted to the site-frequency spectrum (SFS) for putatively neutral exon-flanking sequences for 51 high-coverage Yoruba genomes sequences (Ni: effective population size in epoch i + 1; ti: generations in epoch i + 1). A mixture model for neutral evolution (s = 0), weak negative (s = −1.30 × 10−4), and strong negative (s = −5.86 × 10−4) selection was then fitted to the SFS for coding sequences (CDS) (see Methods). (C) A mixture density network model defines the probabilities of the three components of the mixture model (ℙneutral, ℙweak, ℙstrong) for each possible mutation at each nucleotide site, conditional on the local genomic features. These probabilities allow the likelihood of the polymorphism data to be computed under the Poisson Random Field (PRF) model, using diffusion approximation methods. The parameters of the network are estimated by maximum likelihood, by treating the (negative) log likelihood as a loss function for the neural network. After training, the weights for the three mixture components define a coarse-grained distribution of fitness effects for all potential mutations at each site. This distribution is summarized by a single expected value of |s| for each mutation.











