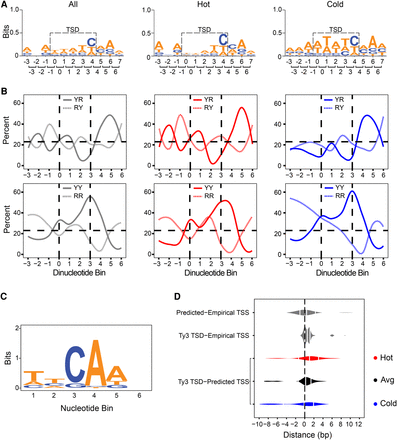
Ty3 insertion site analysis upstream of tDNA genes. (A) WebLogo analysis of the 11 bp comprised of Ty3 5-bp target site duplication (TSD) and ±3 bp flanking sequence. Each TSD was weighted by the total number of Ty3 insertions at that site. Brackets indicate the corresponding nucleotide positions (top row of numbers) assigned to each dinucleotide bin (bottom row of numbers). (B) Plots of dinucleotide frequency determined from sequences shown in A. Dinucleotide starts at position indicated (“0” = dinucleotide at positions 0 and 1 of TSD, etc.). YR/RY (top) and YY/RR (bottom) plots of TSD and flanking sequence shown in A. Vertical dashed lines mark the dinucleotide bins representing the borders of the TSD. Horizontal dashed line represents the random frequency of the YR dinucleotide in the S. cerevisiae genome (23.25%). Random frequency of all dinucleotide bins in the S. cerevisiae genome are 23.25% (YR), 23.25% (RY), 26.71% (YY), and 26.79% (RR). (C) WebLogo analysis of conserved motifs within a 23-bp window upstream of all tDNA genes by MEME Suite. All four DNA nucleotides occur at roughly the same frequency at position 6. (D) Distance analysis of Ty3 TSD to MEME-predicted and empirically determined TSS upstream of tDNAs. From top to bottom: distance between MEME-predicted TSS and TSS of 29 tDNAs empirically determined by Yukawa et al. (2011); distance between Ty3 TSD and empirical TSS; distance between Ty3 TSD and MEME-predicted TSS of all tDNAs in this study categorized by hot, average, and cold phenotypes. For all comparisons, distance is measured from the fifth base of the TSD to the first conserved “C” nucleotide in both MEME-predicted motifs and empirically determined motifs (see text for detailed explanation). The first, second, and third quartiles of each data set are denoted by white lines on each violin plot.











