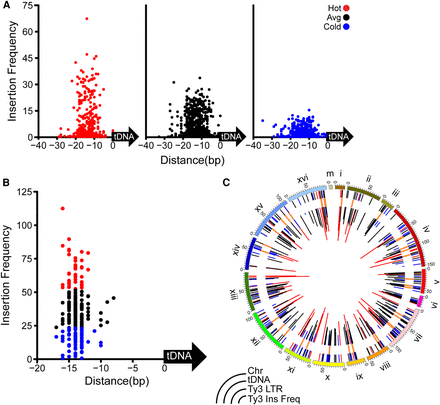
Genome-wide Ty3 insertion mapping using barcode-tagged elements. (A) Ty3 insertion sites are plotted with aligned mature tRNA-coding sequence (tDNA) = 0th bp and insertion site defined as the tDNA proximal end of the 5-bp TSD. Each dot represents the normalized unique barcodes of insertions starting at the strand transfer site proximal to the tDNA per 10,000 hits: (red) hot; (black) average; (blue) cold. Note that classification into hot, average, and cold was based on subsequently aggregating insertions per a 50-bp upstream window of the tDNA. (B) Insertions from A are binned across a 50-bp window upstream of the tDNA (colored as in A). (C) Circos tracks display chromosomes, tDNAs, Ty3 LTRs, and Ty3 insertion frequency: (outside to inside) colored as in A; (Ty3 LTRs) peach.











