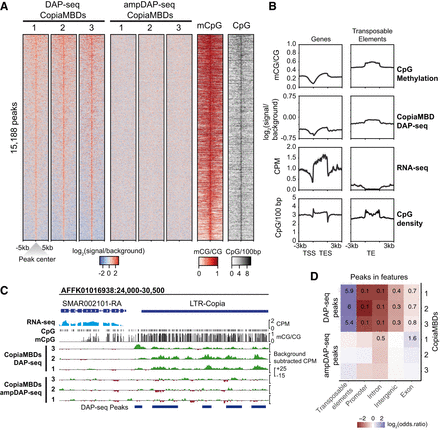
Retrotransposon-encoded MBDs preferentially bind methylated CpG-dense regions overlapping transposable elements. (A) Heatmap showing enrichment levels for three phylogenetically distinct CopiaMBDs based on Strigamia DAP-seq (native genome methylation) and ampDAP-seq (amplified genome depleted of native methylation) libraries, CpG methylation, and CpG density on a union of all three CopiaMBD DAP-seq peaks set. (B) Profiles of CpG methylation, CopiaMBD 3 DAP-seq enrichment, RNA-seq, and CpG density on Strigamia protein-coding genes and transposable elements. (C) Genome browser display showing enrichment tracks for DAP-seq, ampDAP-seq, and cytosine methylation. RNA-seq shown as counts per million (CPM), whole-genome bisulfite sequencing (WGBS) shown as the mC/C ratio at CpG sites, DAP-seq, and ampDAP-seq data shown as background subtracted CPM tracks (CopiaMBD signal minus empty pIX-HALO plasmid). (D) Heatmap showing the enrichment of CopiaMBD peaks on various genomic features. Colors represent log2(odds ratio), whereas untransformed odds ratios are shown for the significantly enriched tests (Fisher's exact test P < 0.01).











