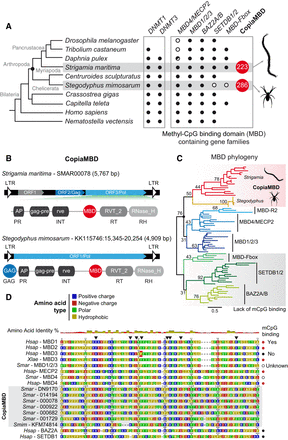
A family of arthropod Copia retrotransposons have incorporated an MBD into their coding sequence. (A) Cladogram showing a subset of animal species, taxonomic affiliation, and the presence/absence patterns of major DNA methylation enzymes (DNMT1 and DNMT3) and MBD gene family members encoded in their genomes. In red indicates the total number of MBDs belonging to Copia retrotransposons encoded in centipede and spider genomes. (B) Case examples of CopiaMBD structure from Strigamia and Stegodyphus genomes, as well as domain architecture of the Pol ORF. The protein domains as defined by Pfam (MBD PF01429, gag_pre PF13976, rve PF00665, RVT_2 PF07727, RNase_H PF00075). The domains as also annotated according to retroviral nomenclature conventions: (GAG) group-specific antigen; (PR) protease; (INT) integrase; (RT) reverse transcriptase; (RH) RNase H. (C) Maximum likelihood phylogeny of the MBD metazoan gene families. Nodal supports represent nonparametric bootstrap as computed by IQ-TREE. Shaded in red are the sequences belonging to Copia retrotransposons, shaded in gray are the MBD families known for not having methyl-binding activity despite encoding an MBD. Red branches indicate CopiaMBD Strigamia sequences, and orange branches indicate CopiaMBD Steodyphus sequences. (D) MBD multisequence alignment. Black triangles highlight amino acids known to influence the DNA binding ability of the MBD (Hendrich and Tweedie 2003). Shaded in red is the phenylalanine of the mammalian MBD3 responsible for its lack of methylcytosine binding activity. Amino acid color code is as per polarity described in the legend. (Hsap) Homo sapiens; (Xlae) Xenopus laevis; (Smar) Strigamia maritima; (Smim) Stegodyphus mimosarum. Silhouettes were obtained from http://phylopic.org/.











