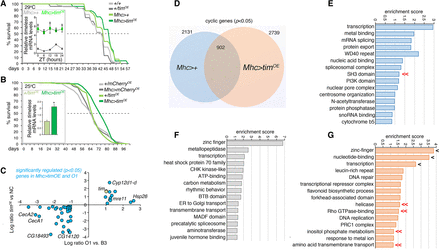
Muscle-specific timeless overexpression extends lifespan and reshapes the muscle circadian transcriptome. (A) Lifespan extension with muscle-specific timeless overexpression (Mhc > timOE) compared to isogenic controls (29°C; P < 0.05; Mhc > timOE n = 70; +/timOE n = 43; Mhc>+ n = 52; +/+ n = 40). (B) Similar results are obtained at 25°C by comparing Mhc > timOE with flies with mCherry overexpression (Mhc > cherryOE) and UAS transgene-alone controls (P < 0.05; Mhc > timOE n = 276; +/timOE n = 255; Mhc > cherryOE n = 381; +/cherryOE n = 315). The level of timeless overexpression obtained with UAS/Gal4 in shown in A,B. (C) Analysis of gene expression changes induced by timeless overexpression reveals that there are some genes significantly and consistently regulated in Mhc > timOE and O flies versus their respective controls (n = 3; genes with P < 0.05 in both genotypes are shown). (D) Comparison of cyclic genes detected in the muscle of Mhc > timOE versus Mhc>+ flies reveals distinct gene sets that are genotype-specific or shared. GO categories describing the cyclic genes that are specific for Mhc>+ are shown (E), as well as the GO categories for genes that cycle in both Mhc>+ and Mhc > timOE (F), and that cycle only in Mhc > timOE (G). Genes that similarly oscillate in O3 versus B3 and Mhc > timOE versus Mhc>+ are identified by double arrows (<<) whereas single arrows (<) highlight GO categories that are shared by multiple gene sets in (E–G).











