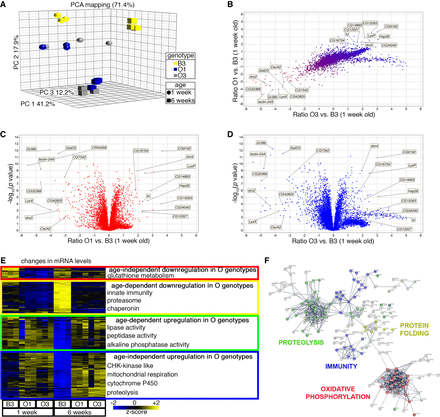
RNA-sequencing identifies gene expression changes associated with protection from skeletal muscle aging. (A,B) PCA mapping identifies clustering of O1 and O3, compared with B3, suggesting that there are genomic variants that affect gene expression consistently in O1 and O3 versus B3, as also indicated by the comparison of the log ratios of O1 versus B3, and O3 versus B3 (B). (C,D) Several genes are differentially regulated, including some already implicated in longevity in Drosophila, such as Hsp26 and takeout. (E) Genes differentially expressed include: genes involved in glutathione transferase activity (red cluster), which are down-regulated in O genotypes; genes involved in innate immunity, secreted factors, the proteasome, and chaperones (yellow cluster), which are induced by aging in the B3 but not in the O lines; genes necessary for proteolysis and peptidase and lipase activity (green cluster), which are down-regulated in B3 with aging; and genes that are up-regulated in O lines at all ages (blue cluster), which include genes necessary for mitochondrial ATP synthesis, oxidative phosphorylation, and cytochrome P450 activity. (F) Similar gene categories are also identified by hub analysis of the most regulated genes that differentiate the O lines from the parental B3.











