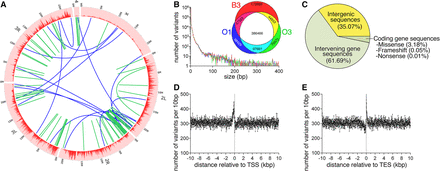
Whole-genome sequencing identifies genetic variants associated with longevity and protection from age-related skeletal muscle functional decay. (A) Circos plot representing the distribution of genomic variants across the euchromatic genome in 20-kb bins. Large intrachromosomal and interchromosomal rearrangements are shown in green and blue, respectively. (B) The distribution of the number of variants by the nucleotide length for the B3, O1, and O3 genotypes relative to the dm3 Drosophila reference genome shows a relatively large proportion of short, in particular single nucleotide polymorphisms. The Venn diagram shows variants that have significantly different allelic frequencies and that are not overlapping between genotypes. Variants unique to the long-lived (O1 and O3) or control (B3) genotypes are shown in cyan and red, respectively, and account for a total of 241,550 variants potentially associated with delayed aging. (C) The proportion of the 241,550 genotype-unique variants and their genomic localization relative to gene sequences. While a large proportion, 64.93%, of total variants are present within a gene sequence, only 3.24% of longevity-associated variants are present within the coding sequence and therefore could potentially affect translation as either missense, nonsense, or frameshift mutations. (D,E) Genotype-unique variants within ±10 kb of either the transcription start site (TSS) and the transcription end site (TES). There is a large increase in variant density immediately prior to the TSS and immediately following the TES.











