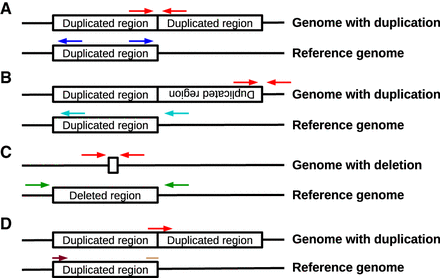
Three types of discordant read pairs (A–C) and breakpoint reads (D) were used to identify different CNV alleles. (A) In tandem duplications, read pairs derived from segments spanning the CNV breakpoint (red arrows) align facing away from each other around the breakpoint on the reference genome (dark blue arrows). (B) In tandem inversions, read pairs derived from segments spanning the end of the inverted segment (red arrows) align facing in the same direction as each other around the breakpoint on the reference genome (cyan arrows). (C) In deletions, read pairs derived from segments spanning the deleted sequence (red arrows) align in the correct orientation around the breakpoint, but farther apart than expected given the insert size of the sequencing library (green arrows). (D) In any of the aforementioned types of CNV (tandem duplication shown here as an example), reads crossing the breakpoint (red arrow) will only partially align on either side of the breakpoint. For the tandem duplication shown here, the start of the read (light brown start of an arrow) aligns at the end of the duplicated region, whereas the end of the read (dark brown end of an arrow) aligns at the start of the duplication.











