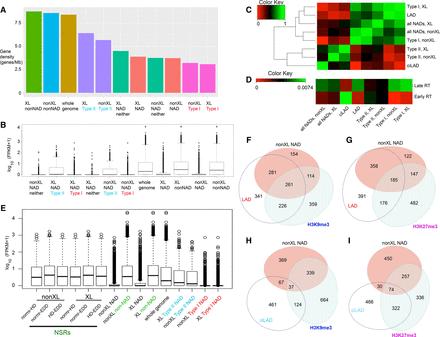
Differential gene density, expression, and chromatin state in different classes of NADs. (A) Gene densities of the indicated genomic regions are shown, ranked left to right. (B) Distribution of gene expression levels from RNA-seq data, expressed as log10(FPKM + 1) for the same genomic regions analyzed in A. (C) Jaccard similarity coefficients were clustered to visualize similarities among the indicated genomic regions. Type I NADs cluster with LADs, Type II NADs cluster with ciLADs. (D) As in C, in this case showing comparison of the indicated regions with MEF early and late RT data (Hiratani et al. 2008, 2010). The scale of this figure is different than C because the RT data is from microarray experiments with much sparser representation than deep sequencing data. (E) As in B, in this case including the NAD-splitting regions (NSRs). NSRs are regions between NADs determined by NADfinder, which fall within large contiguous peaks determined by the indicated two other software programs (for examples, see Fig. 1E; Supplemental Fig. S3B–D). That is, NSRs are uniquely identified by NADfinder as non-NADs in these comparisons. NSRs from the XL and nonXL NAD data sets were analyzed separately. In all cases analyzed, NSRs were found to have FPKM distributions most similar to the non-NAD regions of the genome. (F) Venn diagram analysis of the overlap among NADs (nonXL), LADs, and H3K9me3 peaks. Numbers indicate size of each region (Mb), which are drawn proportional to size. (G) As in F, except that the overlaps with LADs and H3K27me3 peaks are shown. (H) As in F, except that the overlaps with ciLADs and H3K9me3 are shown. (I) As in F, except that the overlaps with ciLADs and H3K27me3 are shown.











