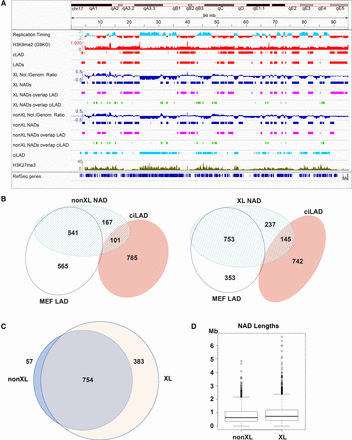
Distinct classes of mouse NADs. (A) Subdivision of MEF NADs based on overlap with LAD or ciLAD regions. Chr 17 is shown. Features associated with heterochromatin (H3K9me2, lamin association, and late RT) are in red, and features associated with euchromatin (early RT and ciLAD regions) are in cyan. At the top is shown DNA RT (Hiratani et al. 2010), followed by H3K9me2 (from MEF cells lacking methyltransferase EHMT2 (also known as G9a), making heterochromatic H3K9me2 more easily visualized) (GSM2803100) (Chen et al. 2018). Next are constitutive LAD and MEF LAD peaks (Peric-Hupkes et al. 2010; Meuleman et al. 2013). Following those are eight tracks of NAD data, the top four from a cross-linked preparation (XL experiment #26), and the bottom four from a non-cross-linked preparation (nonXL experiment #29). At the top of each set is the log2 ratio of nucleolar/genomic sequencing reads (Nol./Genom. Ratio), followed by the peaks determined by NADfinder (blue). These are followed by a pair of tracks in which the NAD peaks were separated at the nucleotide level into regions that overlap the MEF LADs (magenta) and regions that overlap ciLADs (light green). The bottom two tracks are ciLADs in cyan (Peric-Hupkes et al. 2010; Meuleman et al. 2013), and H3K27me3 in olive green (GSM2417097) (Chronis et al. 2017). (B) Venn diagram illustrating the overlap among NADs, LADs, and ciLAD regions. Data from non-cross-linked experiments are on the left, and cross-linked ones are on the right. Numbers show the size of the indicated regions (Mb). (C) Venn diagram illustrating overlap of XL and nonXL NAD peaks. Numbers indicate size of each region in Mb, which are drawn proportional to size. (D) Box plot distributions of NAD lengths, showing median and quartiles in the box.











