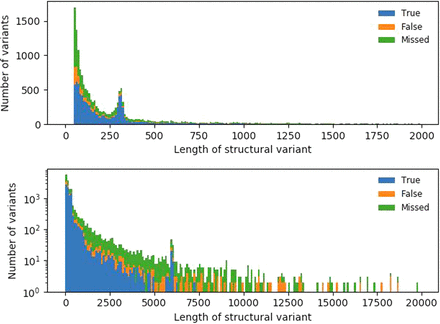
Figure 4.
SV validation status per length SVs identified using Sniffles after NGMLR alignment, compared with truth set. The top panel has SVs up to 2 kb binned per 10 bp; the bottom panel, up to 20 kb, binned per 100 bp with a log-transformed number of variants.











