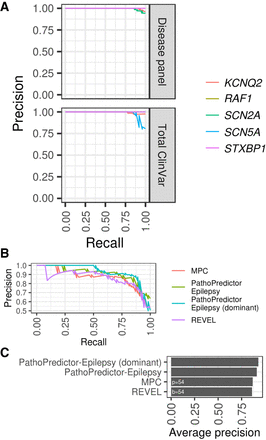
PathoPredictor performance. (A) Precision-recall curves are shown for select genes evaluated during cross-validation with the disease panel data set and tested with ClinVar variants. The curve for RAF1 closely follows and is obscured by that of SCN2A. For KCNQ2, PathoPredictor had an accuracy of 95% for panel variants and 96% for ClinVar variants. (B) PathoPredictor epilepsy-specific classifiers were compared to REVEL and MPC. De novo missense variants in epilepsy panel genes were used as pathogenic variants. Epilepsy panel gene missense variants from unaffected siblings of autism spectrum disorder patients were used as benign variants. PathoPredictor was trained as in Figure 4, but only utilizing the full and dominant epilepsy data sets. Variants were filtered using the same methods applied to ClinVar variants, and additional filters were applied to remove training data for MPC. (C) We summarized each scoring metric's precision-recall curve as the average precision. Both PathoPredictor classifiers achieved a greater average precision than REVEL (P < 0.05), and the dominant epilepsy classifier performed better than MPC (P < 0.05). (p) pathogenic; (b) benign.











