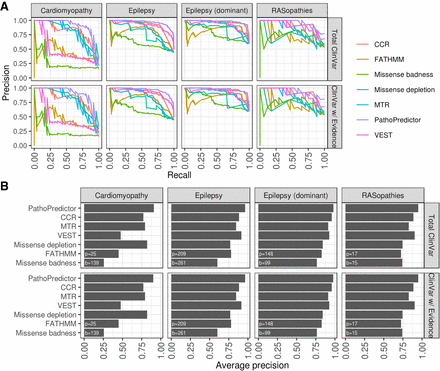
Disease-specific classifier performance using ClinVar data for training and disease panel data for testing. For each disease panel, we collected ClinVar variants in panel genes, using either all ClinVar variants (Total ClinVar) or reviewed ClinVar variants (ClinVar w/Evidence). PathoPredictor training and evaluation for each disease panel proceeded with a hold-one-gene approach. Disease panel variants from the gene of interest were used for evaluation, and ClinVar variants from all remaining disease panel genes were used for training. Using the held-out gene variant prediction scores, we computed a precision-recall curve (A) and summarized the curve with its average precision (B). We then computed a precision-recall curve for each individual feature using untransformed scores. The numbers of pathogenic (p) and benign (b) variants investigated are shown at the bottom left of each panel in B. PathoPredictor performed better than each of its six features (P < 0.05), except for missense depletion for cardiomyopathy panel variants, CCR, missense badness, and VEST for RASopathy panel variants, and CCR for dominant epilepsy panel variants.











