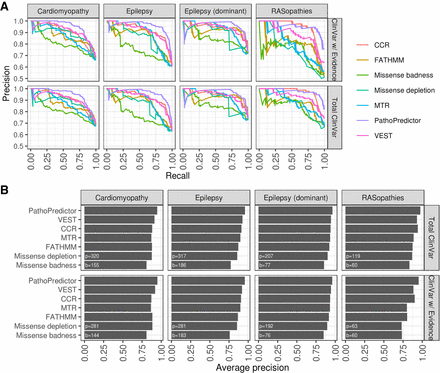
Disease-specific classifier performance using disease panel data for training and ClinVar data for testing. For each disease panel, we applied the hold-one-gene-out models from Figure 3 to ClinVar variants from the held-out gene to obtain pathogenicity prediction scores. We compared PathoPredictor to each feature using a precision-recall curves (A) and average precisions summarizing each curve (B). We used either all ClinVar variants (Total ClinVar) or ClinVar variants with a review status that included at least one submitter or an expert panel (ClinVar w/Evidence). The numbers of pathogenic (p) and benign (b) variants investigated are shown at the bottom left of each panel in B. PathoPredictor performs significantly better than any single feature when examining all ClinVar variants (P < 0.03).











