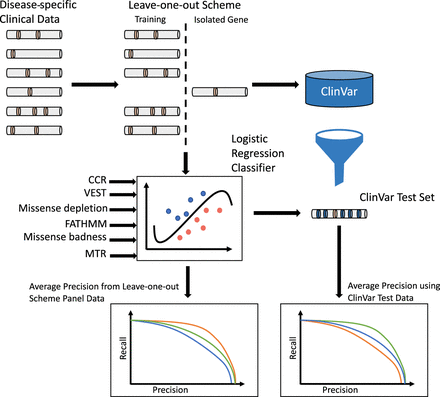
Method description. Our goal was to build disease-specific classifiers of missense variant pathogenicity using variants from clinical panels. For all genes in a disease panel, we trained a model using variants from all other genes except the gene in question and tested the model using variants from that gene of interest. We then used ClinVar variants from the gene of interest as an independent test set. Test results were summarized as average precision scores.











