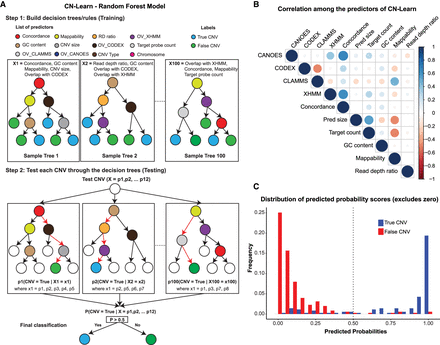
Illustration of the Random Forest model used to build CN-Learn. (A) The inner workings of the Random Forest model used for training CN-Learn is shown. Twelve features were used to grow 100 trees with different subsamples of predictors and training data to classify each CNV in the test set as either true or false. If the predicted probability score was >0.5, the CNV call was classified as true. Calls with predicted probability score <0.5 were labeled as false. (B) A Spearman's rank correlation between pairs of quantitative predictors used by the CN-Learn classifier is shown. The color of the circles indicates the direction of the correlation, and the size of the circles indicates the strength of the correlation. The correlation scores are provided in Supplemental Table 1. (C) The frequency of microarray-validated and -invalidated CNVs, distributed across 20 bins of increasing predicted probability scores, is shown. For the probability bins <0.5, the proportion of CNVs that were validated was higher than the proportion of CNVs that were not validated. This indicated that the classification score of 0.5 is an appropriate threshold for distinguishing true and false CNVs.











