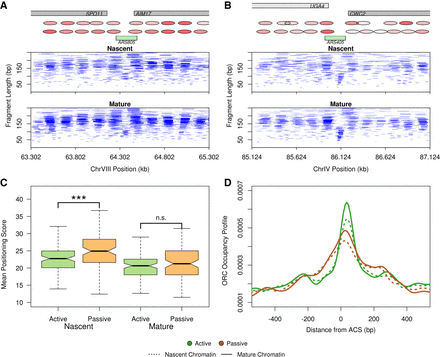
Chromatin maturation at actively and passively replicated origins. (A) Chromatin occupancy profile for nascent and mature chromatin at the active origin ARS805. (B) Chromatin occupancy profile for nascent and mature chromatin at the inactive origin ARS405. Green boxes in A,B represent origins of replication. Gray boxes represent genes on the positive (light gray) and negative (dark gray) strands. A representation of nucleosome positioning and occupancy (intensity) is highlighted with red ovals above the chromatin plots. (C) Passively replicated origins have slower chromatin maturation kinetics. Boxplots depicting the distribution of nucleosome positioning scores for nascent and mature chromatin at 100 active and 100 inactive origins. Nascent chromatin surrounding passively replicated origins is more disorganized than at active origins. t-test: (***) P ≤ 0.001; (n.s.) P > 0.05. (D) ORC occupancy is decreased at passively replicated origins. Density distribution of ORC occupancy footprints as determined from the NCOPs.











