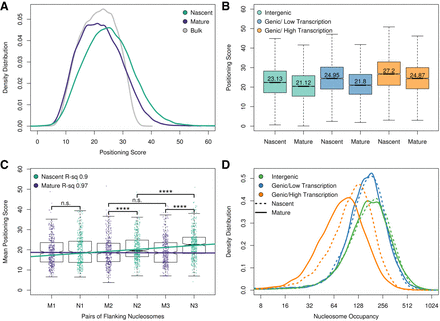
Genome-wide nucleosome positioning and occupancy in nascent and mature chromatin. (A) Genome-wide distribution of nascent, mature, and bulk nucleosome positioning. Approximately 70,000 high-confidence nucleosome dyads were identified in bulk chromatin. For each chromatin fraction (nascent, mature, and bulk), the distance from the midpoint of each sequencing read to the nearest nucleosome dyad was calculated as a nucleosome positioning score. (B) Nascent and mature chromatin organization (nucleosome positioning scores) at intergenic and intragenic nucleosomes. Intragenic nucleosomes were subdivided into high (top 10%) and low (bottom 10%) transcriptional activity. (C) Transcription factors influence chromatin maturation kinetics. Positioning scores of the first, second, and third pairs of nucleosomes flanking 436 transcription factors with strong occupancy in mature chromatin. t-test: (****) P ≤ 0.0001; (n.s.) P > 0.05. (D) Active transcription displaces nucleosomes from mature chromatin. Distribution of nucleosome occupancy scores (sequencing reads assigned to individual nucleosomes) for nascent and mature chromatin at intragenic and intergenic sequences. Actively transcribed genes have more nucleosome occupancy in nascent chromatin than in mature chromatin (P < 2.2 × 10−16).











