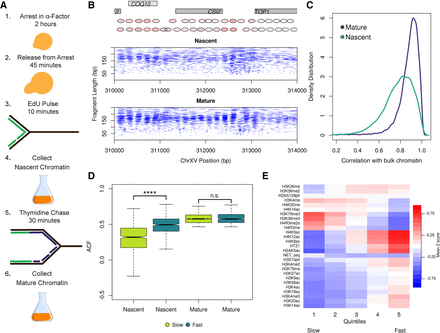
Nascent chromatin occupancy profiles (NCOPs). (A) Schematic of experimental design for capturing EdU-labeled nascent and mature chromatin. (B) NCOPs reveal the maturation of chromatin behind the DNA replication fork at a representative locus, CSI2. Gray boxes represent genes on the positive (light gray) and negative (dark gray) strands. A representation of nucleosome positioning and occupancy (intensity) is highlighted with red ovals above the chromatin plots. (C) Nascent chromatin organization in gene bodies is less organized than mature chromatin. Distribution of correlation scores between nascent and bulk (teal) or mature and bulk chromatin (purple). (D) Differential chromatin maturation kinetics for genes with well-organized mature chromatin. The autocorrelation function (ACF) was used to identify the top 50% of genes with regularly phased arrays of nucleosomes from mature chromatin. These 2700 genes were then binned into quintiles based on the correlation between nascent and mature chromatin. The first and fifth quintiles represent genes with slow and fast chromatin maturation kinetics. The distribution of ACF values (as a proxy for gene organization) in nascent and mature chromatin is depicted for the genes with slow (light green) and fast (dark green) chromatin maturation kinetics. The difference in ACF values among fast and slow maturing chromatin was significant (P < 2.2 × 10−16). (E) Heatmap representing mean Z-score values for 25 histone post-translational modifications, the histone variant H2AZ (encoded by the HTZ1 gene), and NET-seq scores for the individual quintiles of the correlation between nascent and mature chromatin for the 2700 genes described above.











