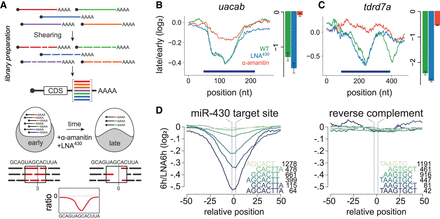
RESA identifies regulatory RNA elements. (A) Schematic of RESA method. Sequence fragments derived from endogenous transcripts are placed within the 3′ UTR of reporter mRNAs. Regulatory activity is inferred from relative depletion or enrichment postinjection. Treatment with α-amanitin or LNA430 delineates sequences under zygotic and/or miR-430 regulatory pathways, respectively. (B,C) RESA identifies sequence regions within uacab and tdrd7a transcripts that are responsive to zygotic regulatory mode. Bar graphs display independent validation by qRT-PCR with reporters containing sequence inserts spanning regulated regions (dark blue bars): late/early fold-change in untreated (green), LNA430-treated (blue), α-amanitin treated (red). (D) Mean destabilization across all loci (minimum coverage of five counts per million at 2 hpf) centered on miR-430 target site variant, with number of represented loci indicated for each variant. Right panel shows mean destabilization for loci possessing the reverse complement for each miR-430 target sequence.











