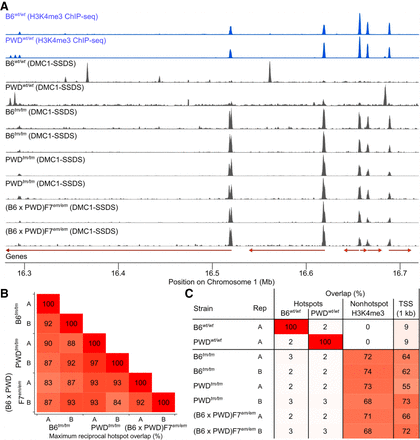
Hotspot locations are consistent in all mice lacking functional PRDM9. (A) DSB hotspots in wild-type mice (B6wt/wt and PWDwt/wt) are absent in mice lacking functional PRDM9 (B6tm/tm, PWDtm/tm, (B6 × PWD)F7em/em). Raw coverage is shown in 150-bp windows. Each panel is scaled to the maximum value. DSB hotspots are the peaks in DMC1-SSDS coverage. Red arrows represent Ensembl gene models. (B) Hotspot locations were conserved in all mice lacking functional PRDM9. The maximum reciprocal hotspot overlap (±500 bp) between pairs of samples is shown. (C) Hotspots in mice lacking functional PRDM9 occur at non-PRDM9-mediated H3K4me3 (non-hotspot) marks and transcription start sites (TSSs). All H3K4me3 peaks (Methods) that did not coincide with a wild-type DSB hotspot were considered non-hotspot H3K4me3. For TSSs, the overlap with Ensembl transcript's 5′ ends ± 500 bp is shown. These features rarely coincide with DSB hotspots in wild-type mice. Note that B6wt/wt H3K4me3 ChIP-seq data are from (Baker et al. 2014). B6wt/wt and PWDwt/wt DMC1-SSDS data are from Smagulova et al. (2016).











