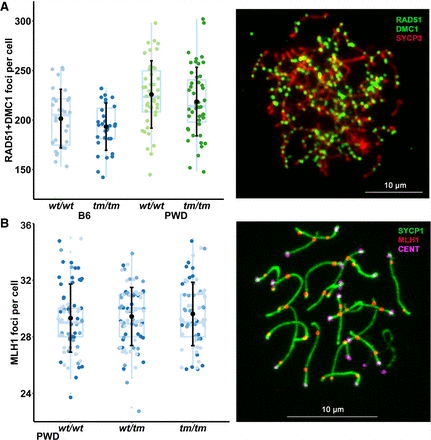
Prdm9 dosage effect on crossover rate and DSB rate in PWD males. (A) PWD males have more DSBs than B6, independent of Prdm9 genotype (P = 0.006 for wild-type cells, P = 0.003 for Prdm9-deficient cells; Linear regression model). DNA meiotic recombinase 1/RAD51 recombinase (DMC1/RAD51) foci mark the sites of meiotic DSBs. The mean numbers of foci per genotype are shown as black dots ± SD. Values for individual cells are depicted as colored dots. These data were obtained from pooled prepuberal testes because these are enriched for zygotene spermatocytes. Loss of functional PRDM9 affects meiotic progression in juveniles similarly to adults, as there were more pachytene spermatocytes in 13-d-old PWDwt/wt (9%) versus PWDtm/tm testes (3%, P = 0.0001, χ2 test). The right panel depicts representative immunolabeling of a zygotene spermatocyte nucleus from PWDwt/tm. (B) The crossover rate (autosomal MLH1 foci per cell) is unchanged in PWDwt/wt and PWDtm/tm mice. The black dots with bars represent the mean ± SD; blue dots depict per-cell values (different shade for each individual male). See Supplemental Table S1 for values. The right panel depicts representative immunolabeling of a pachytene spermatocyte nucleus from PWDwt/tm for synaptonemal complex (SYCP1) and centromeres (CENT).











