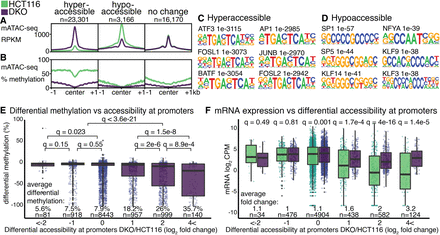
Accessibility and methylation at peaks. (A,B) Significantly changed mATAC-seq hyperaccessible (log2 fold change > 1, q < 0.01, n = 23,310 peaks), hypoaccessible (log2 fold change < −1, q < 0.01, n = 3166 peaks), and unchanged peaks (| log2 fold change | < 1, q > 0.8 n = 16,170). Motifs enriched in hyperaccessible (C) and hypoaccessible (D) sites compared to unchanged sites. (E) DNA methylation changes at promoters binned by accessibility, reported as the change in methylation ratio of DKO cells relative to HCT116 (DKO/HCT116). (F) mRNA expression changes in DKO cells relative to HCT116, reported as log2 CPM, at genes binned by differential accessibility of their promoters as in E. Q-values are for Wilcoxon tests with Benjamini–Hochberg correction (Benjamini and Hochberg 1995).











