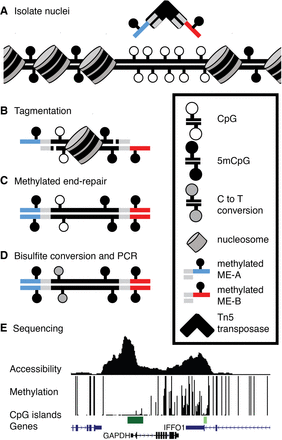
Overview of mATAC-seq. (A) Tn5 carrying methylated oligonucleotides (red and blue segments) is used to (B) perform tagmentation on nuclei at THS sites. (C) Tagmented DNA is end-repaired using 5mdCTP + dDTPs, purified, (D) bisulfite-converted, amplified, and (E) sequenced to measure DNA methylation and accessibility simultaneously. Sample data are shown for one region in HCT116 cells. Peak height in accessibility track is proportional to read abundance; bar height in methylation track is proportional to extent of methylation at CpGs.











