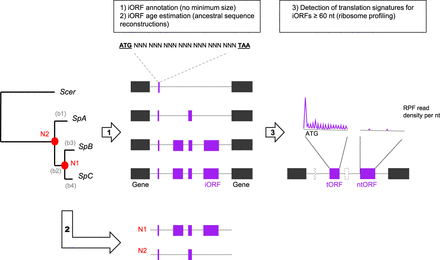
Overview of iORF annotation and translation detection procedure. For a more complete description, see Methods and Supplemental Figure S1. iORF annotation was conducted using S. paradoxus strains that are structured in three main lineages (SpA, SpB, and SpC) with S. cerevisiae as an outgroup. Pairs of genes annotated as syntenic were used to align intergenic genomic regions in which iORFs were characterized. The age of an iORF was estimated using reconstructions of ancestral intergenic sequences at nodes N1 and N2 (in red) to infer their emergence along phylogenetic branches (named b1 to b4, in gray). We chose four strains (one per S. paradoxus lineage and one S. cerevisiae) to characterize the repertoire of translated iORFs (tORFs) using ribosome profiling. iORFs without translation signature were named ntORFs.











