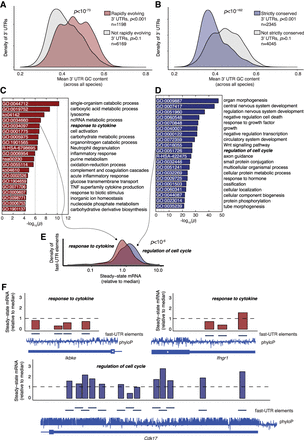
Genes with strictly conserved, AT-rich 3′ UTRs and rapidly evolving, GC-rich 3′ UTRs represent different biological categories. (A) Mean GC content of 3′ UTRs across nine vertebrate species among genes that were found to be rapidly evolving in a multiple sequence alignment versus other genes; (B) the same data as A comparing genes that were strictly conserved in a multiple sequence alignment versus other genes. P-values represent Welch's unequal variance t-test between genes that exhibit strong evidence of conservation/rapid evolution and those that do not. Enriched Gene Ontology categories for genes with rapidly evolving 3′ UTRs (C) or strictly conserved 3′ UTRs (D). P-values are for enrichment of the indicated GO category computed by Metascape. (E) Steady-state mRNA of fast-UTR reporters derived from genes enriched in “response to cytokine” or “regulation of cell cycle” categories with rapidly evolving or strictly conserved 3′ UTRs, respectively. P-value represents Welch's two-sample t-test. (F) Steady-state mRNA of fast-UTR reporters for indicated RBP-occupied regions (labeled “fast-UTR elements”) with conservation across 60 placental mammal species (phyloP) displayed at each nucleotide position.











