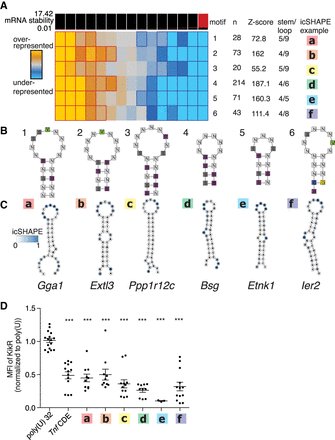
TEISER identifies in vivo–folded 3′ UTR structural motifs that inhibit gene expression. (A) TEISER analysis identifies structural motifs enriched in destabilizing sequences. Columns show enrichment of motifs in deciles of GCLiPP peaks arranged by fast-UTR steady-state mRNA abundance; rows represent individual motifs. Generic motif structures (B) and a predicted structure (C) for an example of each motif is depicted with icSHAPE signal indicated by color. (D) TEISER identified motifs’ lower gene expression. Kikume fluorescent protein synthesis in primary mouse T cells transfected with in vitro–transcribed mRNAs with the indicated sequence inserted downstream from the stop codon. Data represent transfections of a single construct into the T cells from a single mouse pooled from 1–4 experiments using three mice each, with mean and standard error of the mean indicated by line and error bars, respectively. (***) P < 0.0001 in unpaired t-test relative to poly(U).











