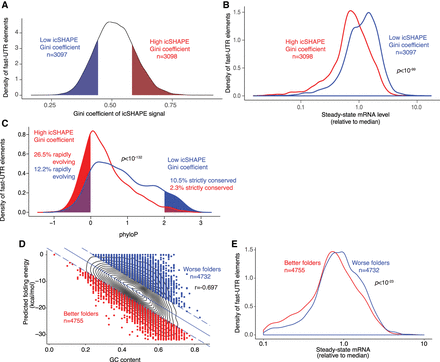
In vivo–folded structures are enriched for gene regulatory activity and are rapidly evolving. (A) RBP-occupied sequences in fast-UTR library with corresponding icSHAPE data were stratified quintiles of icSHAPE Gini coefficient. Strongly structured sequences (red, top quintile icSHAPE Gini coefficient) or nonstructured sequences (blue, bottom quintile icSHAPE Gini coefficient) were identified. (B) Steady-state mRNA abundance in fast-UTR assay for sequences of classes depicted in A. P-value from Welch's two-sample t-test. (C) Sequence conservation across placental mammals for sequences of classes depicted in A. P-value from Welch's two-sample t-test. Shading depicts fraction of sequences in each class (structured or nonstructured) corresponding to rapid evolution (mean phylo P < 0) or strict conservation (mean phylo P > 2). (D) Correlation between GC content and predicted folding energy for RBP-occupied sequences included in fast-UTR library. Plot shows linear regression and top/bottom outliers (∼15% of data points furthest from regression line). (r) Pearson correlation coefficient. (E) Steady-state mRNA abundance in mouse T cell fast-UTR assay for outliers of folding that are better or worse than regression. P-value represents Welch's unequal variance t-test.











