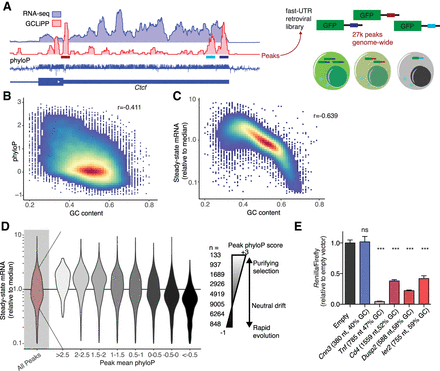
GC-rich 3′ UTR sequences are rapidly evolving and destabilize reporter mRNAs. (A) Schematic diagram of fast-UTR assay. Approximately 27,000 70-nt sequences corresponding to RBP-occupied regions of mouse T cell 3′ UTRs were synthesized and cloned into the 3′ UTR of a murine stem cell virus based GFP reporter. The library was transduced into primary T cells, library inserts amplified from cDNA, and genomic DNA, and gene expression quantified for each insert by calculating a median normalized RNA/DNA ratio. (B) Relationship between GC content and conservation for RBP-occupied sequences. (r) Pearson correlation coefficient. (C) Relationship between GC content and steady-state mRNA abundance in fast-UTR reporter assay for RBP-occupied sequences. (D) Steady-state mRNA abundance in fast-UTR of RBP-occupied sequences, binned on placental mammal conservation. (E) Dual-luciferase assay showing Renilla luciferase activity relative to Firefly luciferase activity, in Th2 cells transfected with plasmid with indicated 3′ UTR downstream from Renilla luciferase gene and control Firefly luciferase gene. A representative experiment using Th2 cell cultures from four mice is shown. Mean and standard error of the mean are indicated by bar graph and error bars, respectively. (***) P < 0.0001 in unpaired t-test relative to empty vector.











