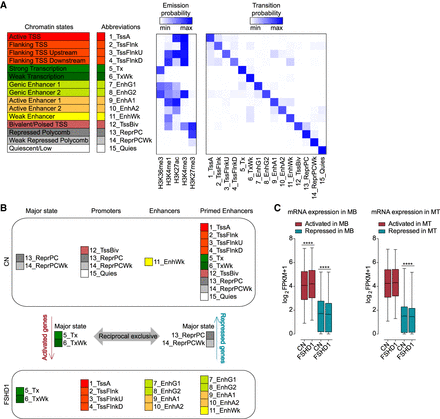
Chromatin segmentation analysis revealed chromatin state switches consistent with transcriptional changes in FSHD1 muscle cells. (A) ChromHMM 15-state model obtained with ChIP-seq data sets for H3K36me3, H3K4me1, H3K27ac, H3K4me3, and H3K27me3. Heatmaps display histone marks emission probabilities and transition probabilities between chromatin states. (B) Schematic representation of the strategy used to assign genes as activated or repressed in FSHD1. (C) Expression levels from RNA-seq data sets for FSHD1 activated and repressed genes in MBs (left) and MTs (right) in CN (CN-3, CN-4, and Yao's data sets C20, C21, C22) and FSHD1 (FSHD1-3, FSHD1-4, and Yao's data sets F4, F6) (Yao et al. 2014). Box-and-whisker plots show the median of matched expression values of each gene for CN and FSHD1, and whiskers extend to the 5 to 95 percentiles. P-values were calculated by paired two-tailed Wilcoxon matched-pair signed-rank test with a confidence interval of 99%. Asterisks represent statistical P-values: for CN versus FSHD1 activated in MBs, P < 0.0001; for CN versus FSHD1 repressed in MBs, P < 0.0001; for CN versus FSHD1 repressed in MTs, P < 0.0001.











