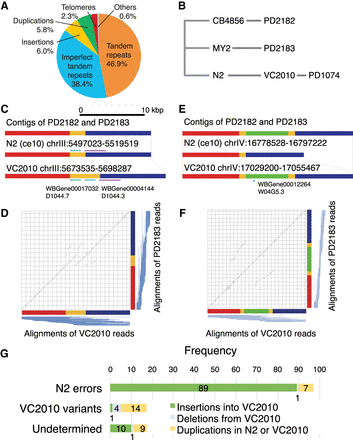
New genomic regions in VC2010 assembly. (A) Subdivision of sequence classes causing the 1.8-Mb increase in genome size from N2 assembly to VC2010. Large tandem repeat expansions (of size >1 kb) are predominant, accounting for 85% of the increased VC2010 DNA. Other sequence classes include insertions (>100 nt), duplications (>100 nt), and telomere repeats. Tandem repeats are divided into some with clear repeat units and others (“imperfect”) without them (Supplemental Fig. S7). (B) Phylogenetic tree of N2, VC2010 (PD1074), and outgroup strains CB4856 (PD2182) and MY2 (PD2183). (C) The yellow-colored duplicated region with two copies of a gene in VC2010 is compared with its best matching regions in N2, PD2182, and PD2183. The comparison implies that the duplication was a recent event occurring in the lineage from the original N2 strain to VC2010. Of note, two duplicated regions overlap slightly. (D) Because long reads were unavailable for N2, we compare the regions in VC2010 and PD2183 for which long reads were available, and we show a dot plot between the regions (a similar dot plot between VC2010 and PD2182 is shown in Supplemental Fig. S12). To confirm the correctness of both regions, we align raw PacBio reads collected from VC2010 and PD2183 to their respective genomic regions, and the alignments are shown as blue lines below the x-axis and to the right of the y-axis. Indeed, a number of alignments span and validate the focal duplicated region and its matching region. (E) A comparison of regions where VC2010, PD2182, and PD2183 coincide, but the green-colored region is missing in the N2 reference assembly, implying that the segment had been lost in culturing animals or clones used for the N2 assembly or in the original N2 assembly process. (F) As in D, aligning raw PacBio reads to both regions in VC2010 and PD2183 shows their validity (a similar dot plot between VC2010 and PD2182 is shown in Supplemental Fig. S10). (G) Frequencies of apparent insertions into VC2010 (missing in N2), deletions from VC2010 (surplus in N2), and genome duplications (in N2 or VC2010), sorted into three categories: 97 assembly errors in the N2 genome, 19 variants that arose in the lineage from N2 to VC2010, and 20 undetermined cases because of inconsistency among the four genomes. We categorized individual large variants by inspecting the dot plots in Supplemental Figures S10–S12 (Supplemental Tables S16–S18). Of the 97 assembly errors, 89 (92%) were regions missing in the N2 reference assembly.











