Figure 1.
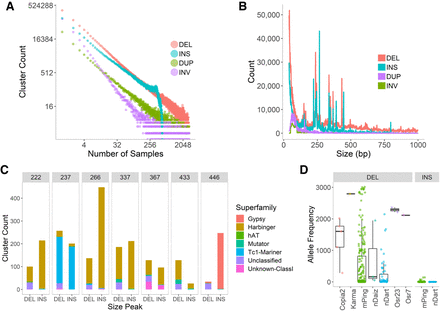
Distribution and classification of SVs. (A) Frequency of observations per SV cluster. Only 562 high-coverage samples were used for insertion detection. (B) Distribution of variant sizes by SV type. (C) Classification of variants in each peak (cluster frequency > 10 samples). (D) Frequencies of events with 98% sequence identity to known or potentially active TEs in rice.











