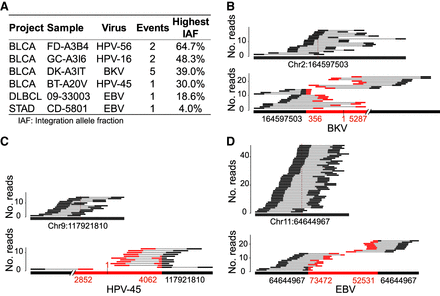
Identifying oncovirus candidates with integrations in bladder cancer, diffuse large B-cell lymphoma, and gastric adenocarcinoma samples. (A) Summary of identified integrations: The Cancer Genome Atlas (TCGA) Urothelial Bladder Carcinoma (BLCA); TCGA Stomach Adenocarcinoma (STAD); The Cancer Genome Characterization Initiative Diffuse Large B-Cell Lymphoma (DLBCL). (B) Sequence read alignment of a BKV (AB485698.1) integration event with 39% integration allele fraction detected in a bladder cancer sample TCGA-DK-A3IT. A total of 23 supporting read pairs, including 19 chimeric and four split reads, were found crossing the two breakpoints, supporting an integration, and 18 read pairs that support no integration were fully mapped to the human reference genome. (C) Sequence read alignment of an HPV-45 (EF202163.1) integration event with 30% integration allele fraction detected in a bladder cancer sample TCGA-BT-A20V. A total of 12 supporting read pairs, including 11 chimeric reads and one split read, were found crossing the two breakpoints, supporting an integration, and 14 read pairs were fully mapped to the human reference genome, supporting no integration. (D) Sequence read alignment of an EBV (AB828191.1) integration event with 18.6% integration allele fraction detected in a diffuse large B-cell lymphoma sample 09-33003. A total of 22 supporting read pairs, including 18 chimeric and four split reads, were found crossing the two breakpoints, supporting an integration, and 48 read pairs that support no integration were fully mapped to the human reference genome.











