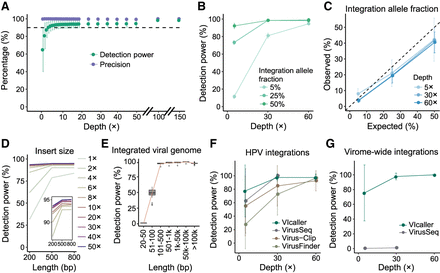
Applying VIcaller to simulation data sets. (A) Detection power and precision were measured by simulated (germline) viral integrations with depths from 1× to 150×. (B) Detection power for integrations with 5%, 25%, and 50% integration allele fractions. On average, 86 viral integrations were used for the calculation. (C) Accuracy of calculated integration allele fractions. (D) Relationship between detection power and insert sizes for paired-end sequence reads at different sequencing depths. (E) Relationship between detection power and lengths of integrated viral sequences. The viral integrations detected under different sequencing depths were combined for the calculation. Comparison of the detection power of VIcaller with existing tools for detecting 10 simulated HPV integrations (F) and 90 simulated virome-wide integrations (G). VirusSeq was only capable of detecting less than 20 human viruses; thus, the detection power was extremely low. It also ran out of server wall time at 60× sequencing depth. VirusFinder and Virus-Clip were not applicable for analyzing data containing the virome-wide integrations.











