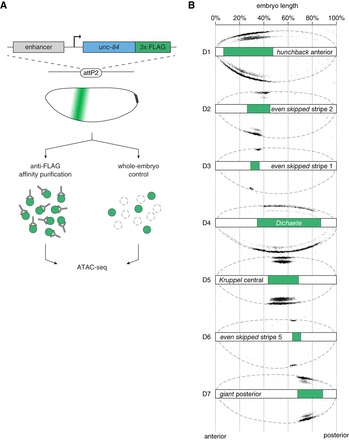
Domain-restricted ATAC-seq profiling along antero-posterior axis in the blastoderm embryo. (A) Selected AP domains are targeted by expressing a nuclear tag, UNC84-3×FLAG, under control of well-characterized enhancers of gap and pair-rule genes. All reporter constructs are integrated at the same genomic site (attP2) (Pfeiffer et al. 2008) to standardize genetic background. After homogenization of staged embryos (cellularizing blastoderm, stage 5, 2:50–3:10 h after egg laying), tagged nuclei are affinity-purified with anti-FLAG antibodies, followed by Tn5 transposase fragmentation and ATAC-seq library preparation. An ATAC-seq library representing an entire pool of nuclei from homogenized embryos (whole-embryo) serves as a control. (B) Overview of the tagged domains (D1–D7). Selected embryos immunolabeled with an anti-FLAG antibody show spatially restricted expression domains of the nuclear tag. Each domain is additionally schematized (green bars) to indicate its position along the AP axis. Embryos are positioned with anterior to the left and dorsal side up.











