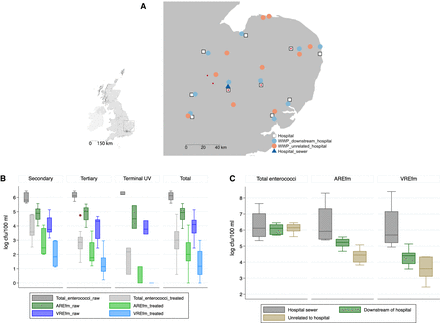
Geographic origin of E. faecium isolates and wastewater counts. (A) Map of 20 wastewater treatment plants in the East of England tested for E. faecium, 10 of which were directly downstream from acute hospitals belonging to the Cambridge University Hospitals NHS Foundation Trust (CUH) referral network. (WWP) Wastewater plant. Red dots indicate the source of bloodstream isolates. (Inset) Map of the United Kingdom, with square denoting the East of England. (B) Log10 counts of E. faecium recovered from 100 mL of wastewater under increasing antibiotic selection for untreated and treated wastewater according to water treatment type: secondary (n = 7), tertiary (n = 10), UV light treatment (n = 3), and for all plants (n = 20). All comparisons between treated and untreated wastewater counts were statistically significant (P < 0.001), as were comparisons of count reductions according to treatment type for each of total enterococci, AREfm, and VREfm (P = 0.02, P < 0.001, and P = 0.001, respectively). (C) Median log10 counts of E. faecium recovered from 100 mL of wastewater under increasing antibiotic selection from untreated wastewater from the main CUH sewer on four separate occasions, from 10 wastewater plants located downstream from acute hospitals, and from 10 wastewater plants unrelated to acute hospital waste. Boxes represent interquartile ranges, whiskers 1.5 times the interquartile range, and dots outside values (B,C).











