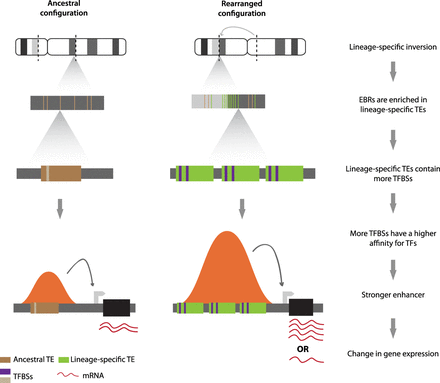
A model for the evolution of chromosome rearrangements with gene expression divergence by means of lineage-specific transposable elements (TEs). Chromosome rearrangement boundaries (EBRs) are enriched for lineage-specific TEs. These TEs harbor a higher number of TFBSs than ancestral TEs; therefore, they have a higher affinity for TFs and a stronger influence in gene expression and regulation than those found elsewhere in the genome. This leads to a higher differential expression for orthologous genes between species with and without the gross genomic rearrangement. Brown and green boxes represent ancestral or lineage-specific TEs, respectively. Purple bars represent TFBSs, and black boxes represent genes. Orange bell-shaped curves represent peaks of H3K27ac as functional enhancers, with the height of the bell proportional to the strength of the enhancer.











