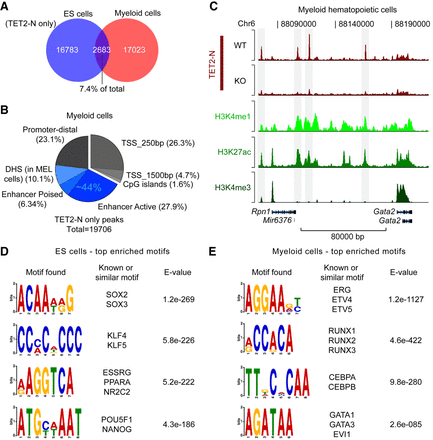
Cell-type–specific binding pattern of TET2 in myeloid hematopoietic cells. (A) Venn diagram showing overlap of TET2 binding sites in ES cells and myeloid hematopoietic cells. Binding sites are defined from biological replicate samples by enrichment over nonspecific ChIP-enriched regions in Tet2 knockout cells using the TET2-N antibody. (B) Pie chart showing distribution of TET2-bound regions in myeloid hematopoietic cells. Each TET2 binding site is counted only once and excluded when moving clockwise from proximal promoters (e.g., 10.1% of TET2 BS overlap with DHS but does not overlap with any of the preceding genomic elements). Blue hues indicate fraction (∼44%) of TET2-bound regions with regulatory potential at promoter-distal sites based on H3K27 acetylation and H3K4 methylation in myeloid cells as well as DHS in the related mouse erythroleukemia (MEL) cell line. (C) Representative ChIP-seq tracks showing TET2-bound enhancer regions upstream of the Gata2 gene in myeloid hematopoietic cells. (D) List of top enriched logos and their associated DNA-binding TFs identified in promoter-distal (−1.5 kb/+500 bp from TSS) TET2-bound regions in ES cells. (E) As in D, but for myeloid hematopoietic cells. Regions in B and C enriched for H3K27ac, H3K4me3, and H3K4me1 were experimentally determined in Rasmussen et al. (2015), and DHS in MEL cells was downloaded from the ENCODE Project (The ENCODE Project Consortium 2011).











