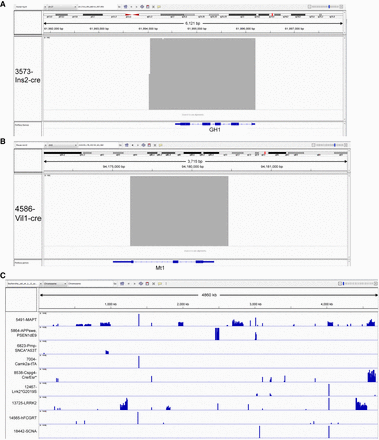
TLA reveals additional passenger cassettes and fragments in transgenes. (A,B) View of TLA reads (indicated in gray above the gene model) that map to the human growth hormone (GH1, also known as hGH) and the mouse metallothionein (Mt1) gene for two transgenes (Ins2-cre and Vil-cre, respectively), showing the inclusion of the entire gene structure, including coding exons. (C) Reads for nine transgenes mapped to the Escherichia coli genome indicating a variable level of coinsertion into the transgene integration site. Deep coverage for discrete loci shared between multiple lines indicates sequences that are part of the transgene vector. The amount of E. coli cointegration ranges from a few hundred bp to more than 200 kb. Short names for each transgene are used for readability and are defined in Supplemental Table S1.











