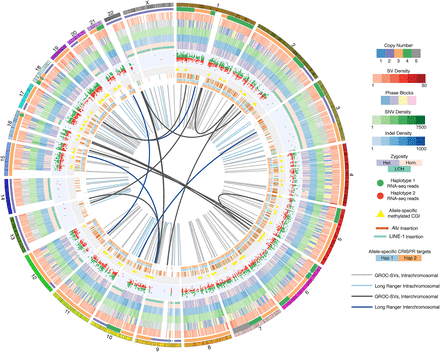
Comprehensive overview of the K562 genome. Circos (Krzywinski et al. 2009) visualization of the K562 genome with the following tracks in inward concentric order: chromosomes; CN, i.e., ploidy by chromosome segment; merged SV density in 1.5-Mb windows of deletions, duplications, and inversions identified using ARC-SV (Arthur et al. 2018), BreakDancer (Chen et al. 2009), BreakSeq (Lam et al. 2010), LUMPY (Layer et al. 2014), Pindel (Ye et al. 2009), and Long Ranger (Zheng et al. 2016; Marks et al. 2018); phased haplotype blocks (demarcated with four colors for clearer visualization); SNV density in 1-Mb windows; indel density in 1-Mb windows; dominant zygosity in 1-Mb windows (heterozygous or homozygous >50%) with regions exhibiting loss of heterozygosity (LOH) indicated; RNA-seq reads for loci exhibiting allele-specific expression; CpG islands (CGI) exhibiting allele-specific methylation; histogram (log-scale) of allele-specifically methylated CpGs in 50-kb windows; nonreference Alu and LINE-1 insertions; allele-specific CRISPR target sites; large-scale rearrangements detected by Long Ranger (Zheng et al. 2016; Marks et al. 2018) (light blue: intrachromosomal; dark blue: interchromosomal); and by GROC-SVs (Spies et al. 2017) (light gray: intrachromsomal; dark gray: interchromosomal).











