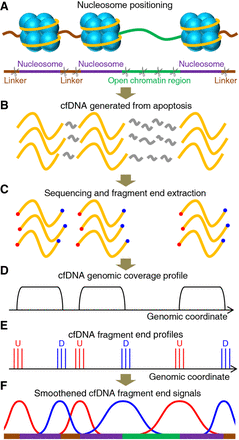
Conceptual framework of cell-free DNA (cfDNA) fragmentation analysis. (A) Illustration of nucleosomes with wrapped DNA (yellow line), linkers (brown line), and open chromatin regions (green line). An abstraction of nucleosome positioning and illustration of cutting events (scissors) during apoptosis were also provided. (B) Illustration of cfDNA generated from apoptotic DNA fragmentation. DNA wrapped around the nucleosomes is preserved, and that in the linkers and open chromatin regions will be cleaved into small pieces (gray line) that cannot be sequenced efficiently. (C) Illustration of the sequenced reads and extraction of the ends. Red and blue represent upstream (U) and downstream (D) fragment ends, respectively. (D) Genomic coverage; (E) U and D end profiles of cfDNA in relation to the reference genome. (F) Smoothened cfDNA end signals and deduced nucleosome positioning. Purple, brown, and green lines represent nucleosomes, linkers, and open chromatin regions, respectively.











