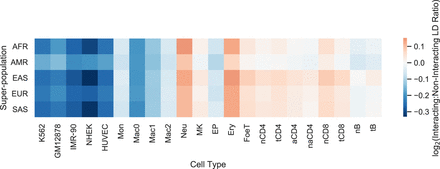
The maximum pairwise LD between SNPs located on the fragments of statistically significant and distance-matched nonsignificant chromatin interactions (interaction LD) was computed for five Hi-C and 17 PCHi-C data sets. The log ratio of mean interaction LD for significant versus nonsignificant interactions quantifies how well LD acts as a proxy for chromatin interactions; a log ratio greater than 0 indicates significant interactions are enriched for SNPs in strong LD. However, the log ratio is near 0 for all cell types and super-populations, indicating that LD is not a sufficient proxy for chromatin interactions. Supplemental Table S2 provides numeric values for this figure; log ratios smaller or larger than 0 are the result of relatively small differences in weak LD.











