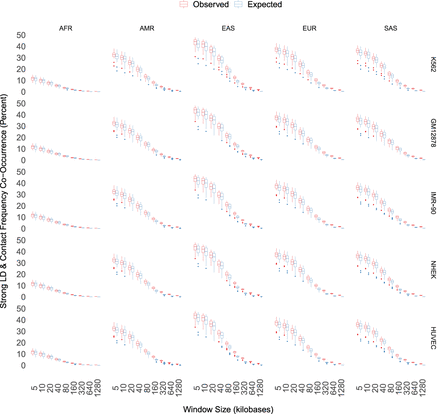
Frequent Hi-C contacts (top 25%) and strong LD (R2 > 0.8) co-occur <50% of the time at short genomic distances. Concordance is cut nearly in half at 40 kb, where most LD has decayed to 0, and is nearly 0 at many scales where statistically significant chromatin interactions occur. Maximum concordance and rate of decay vary by super-population, with AFR having only ∼12% concordance at short genomic distances. LD is elevated compared to expectation at the longest genomic distances, although the effect size is small and median LD is close to zero.











