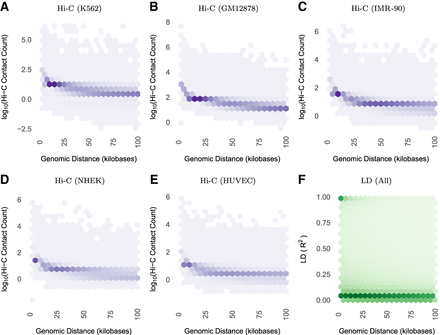
Both (A–E) Hi-C contact frequency (Rao et al. 2014) and (F) LD are anti-correlated with genomic distance (Spearman's correlation between −0.50 and −0.71 for Hi-C across cell lines; it is −0.52 for LD). LD decays toward zero at much shorter genomic distances than contact frequency, with high LD SNP pairs concentrated below 50 kb. In contrast, Hi-C contacts are common up to and exceeding the median length of contact domains (250 kb) or TADs (840 kb). All plots display nonzero values from their respective data sets. Contact frequencies (panels A–E) vary in approximate proportion to sequencing depth and number of replicates per cell line (Supplemental Table S1). Panel F includes 836 million bi-allelic SNP pairs on Chromosome 14, which is representative of other chromosomes. Supplemental Figure S3 shows decay up to 2 Mb, while this figure highlights decay up to 100 kb. Supplemental Figure S4 shows that there is nearly identical LD scaling per super-population.











