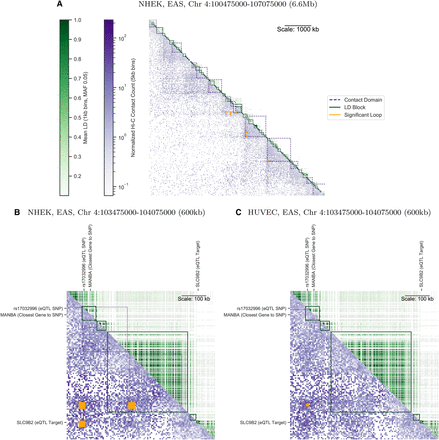
An annotated matrix illustrates differences between the genomic scales of LD (The 1000 Genomes Project Consortium 2015) (R2, upper triangle, green) versus Hi-C contact frequency (Rao et al. 2014) (lower triangle, purple). Rows and columns are binned genomic coordinates (hg19 assembly) with lower bins near the upper left; for example, row 10 column 11 stores the LD between a bin and its neighbor, while row 11 column 10 stores the contact frequency for the same pair of bins. More frequent contacts (5-kb bins) are darker purple; higher LD (averaged over nonzero LD pairs in 1-kb bins) are darker green. Contact domains (nested purple squares) and significant interactions (orange squares) were computed from Hi-C data. LD blocks (green squares) were computed from 1000 Genomes genotypes. While some LD blocks fall within contact domains, there are also many cases where they overlap domain boundaries. (A) A representative 6.6-Mb locus on Chromosome 4 shows Hi-C contacts (NHEK cells) span much longer distances than LD (EAS super-population). (B) A 600-kb locus on the same chromosome illustrates the complexities of mapping a noncoding SNP (rs17032996) to a target gene. The closest gene MANBA falls within the same LD block as the SNP. However, Hi-C data shows the SNP contacts the SLC9B2 gene ∼460 kb away in NHEK cells, skipping over intervening expressed gene MANBA. rs17032996 is also an eQTL in B cells (Fairfax et al. 2012) and significantly interacts with SLC9B2 in several blood cell types (Javierre et al. 2016). (C) In HUVEC cells, the SNP no longer interacts with SLC9B2, and several contact domains are lost.











